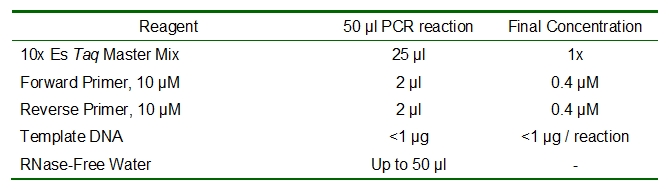
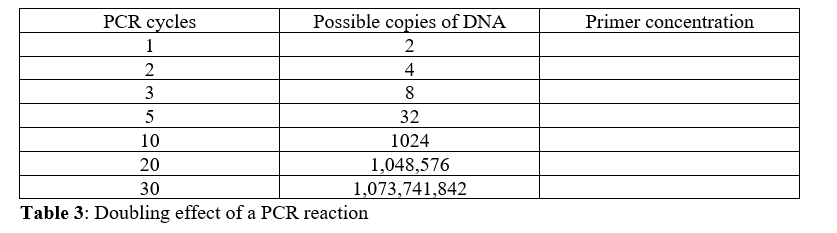
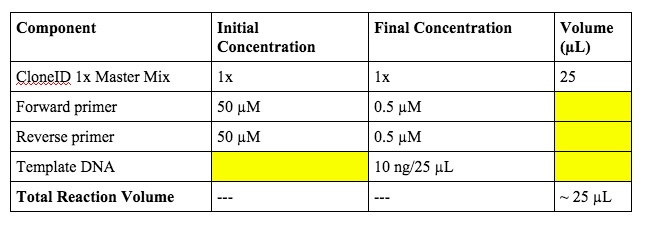
SOLVED: 1. 2.5 μl of 10 μM working primer concentration is added to a 47.5 μl PCR reaction. What is the final concentration of primer in the PCR reaction? 2. Is this

Use of dual priming oligonucleotide system-based multiplex RT-PCR assay to detect five diarrhea viruses in pig herds in South China | AMB Express | Full Text
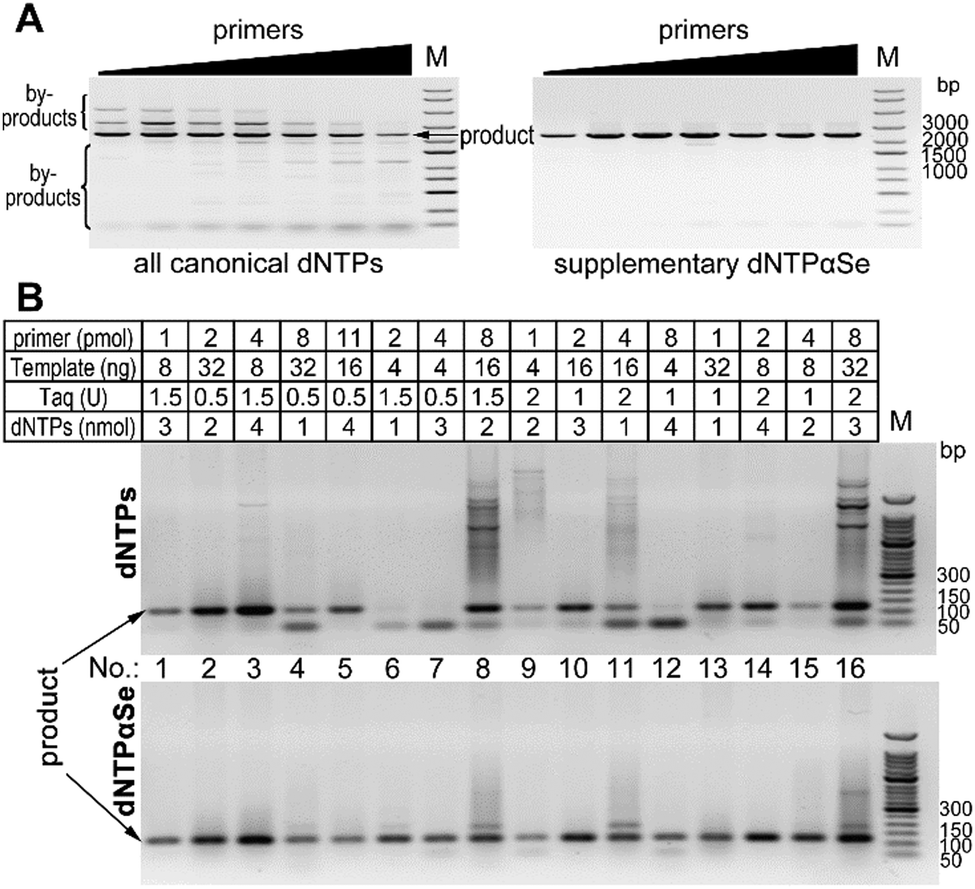
Highly convenient and highly specific-and-sensitive PCR using Se-atom modified dNTPs - Chemical Communications (RSC Publishing) DOI:10.1039/D0CC06172G

Unreacted Labeled PCR Primers Inhibit the Signal in a Nucleic Acid Lateral Flow Assay as Elucidated by a Transport Reaction Model | ACS Measurement Science Au

Figure 2 from COMplementary Primer ASymmetric PCR (COMPAS-PCR) Applied to the Identification of Salmo salar, Salmo trutta and Their Hybrids | Semantic Scholar

Optimization of primer concentration for Cyt b , 12S and COI genes.... | Download Scientific Diagram
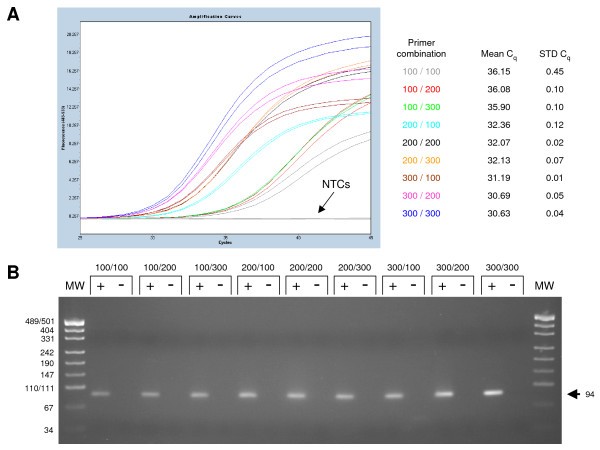
Validation of a primer optimisation matrix to improve the performance of reverse transcription – quantitative real-time PCR assays | BMC Research Notes | Full Text

An optimized protocol for stepwise optimization of real-time RT-PCR analysis | Horticulture Research

SOLVED: Primers: The stock solution of each primer is at a concentration of 100 μM. You are asked to create a new dilution for each of your primers, mixing 1 μl of